Survey
Collection of Scientific Knowledge Graphs (SciKGs) resources
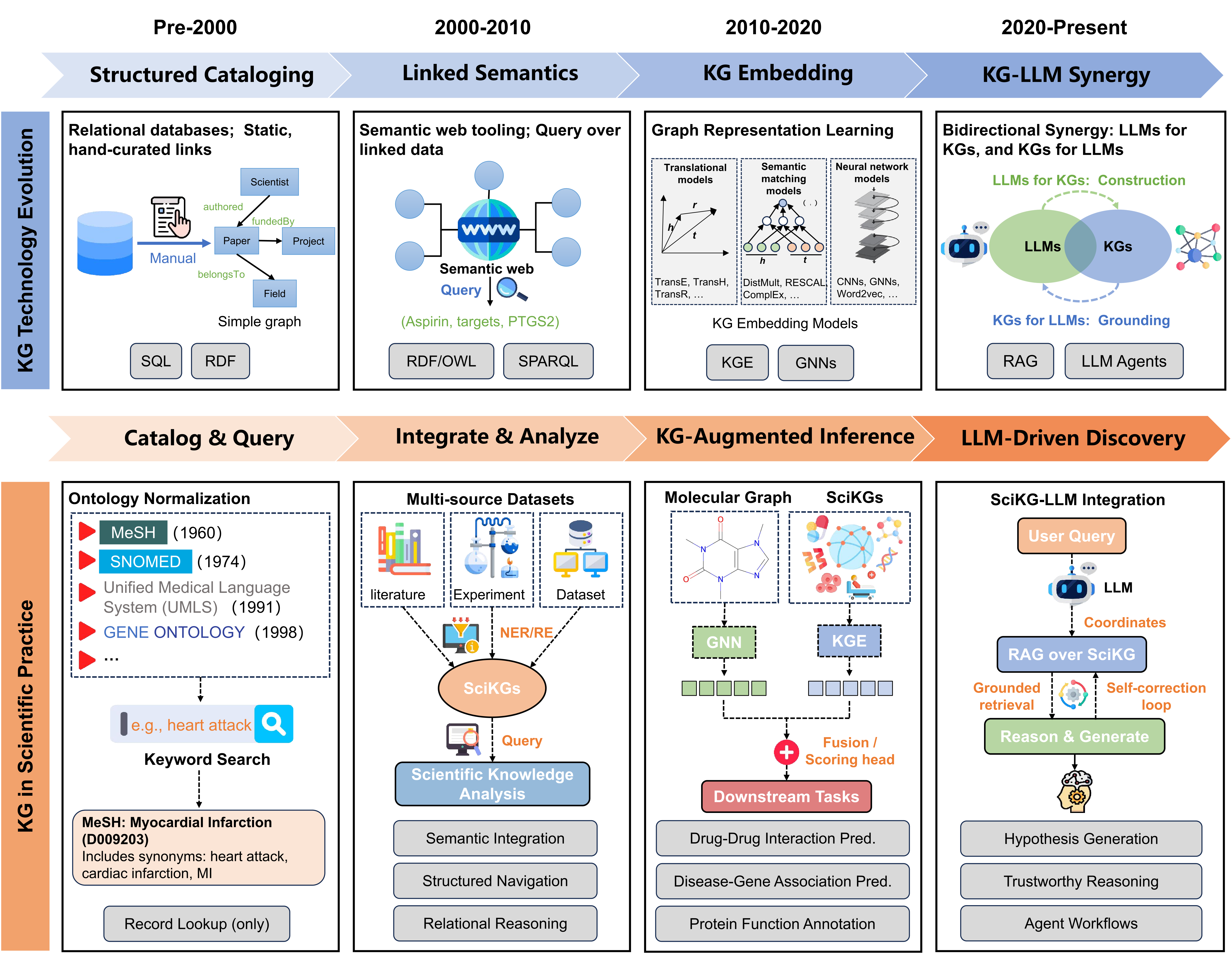
Understand SciKGs: The Backbone of Al for Science
SciKGs reshape research paradigms by converting multi-source scientific knowledge into semantic networks, enabling machine reasoning for tasks like drug discovery and material design.
Leverage Our Curated Resource Hub
Built on a definitive survey, our open GitHub-hosted navigation station provides access to key databases, tools, and applications with direct links and brief explanations.
Accelerate Discovery with SciKG-LLM Collaboration
We deconstruct the emerging ScikG-LLM synergy paradigm to automate scientific discovery, lowering entry barriers and empowering researchers to drive Al-powered progress.
SciKG Resource Hub
SciKG commonly used public databases
authoritative structured data sources covering the fields of biology, chemistry, and materials, which are the cornerstone of building high-quality SciKG.
ScIKG construction and maintenance tool
a software and platform used for tasks such as knowledge extraction, ontology alignment, graph fusion, dynamic updates, and inference completion.
SciKG core application scenario
Focusing on four major application scenarios: drug development and optimization, omics interpretation and analysis, chemical reactions and synthesis, and material design and synthesis.
SciKG-LLM collaborative resources
frameworks, models, and application cases that support bidirectional empowerment of both (such as knowledge grounding, semantic reasoning, hypothesis generation).
Drug Development and Optimization
| Year | Title | KG Name | KG Type | Domain | Construction Method | Venue | Paper |
|---|---|---|---|---|---|---|---|
| 2025 | TarIKGC: A Target Identification Tool Using Semantics-Enhanced Knowledge Graph Completion with Application to CDK2 Inhibitor Discovery | biological activity KG | public KG | DTI prediction | Semi-automated | Journal of Medicinal Chemistry | Link |
| 2025 | A comprehensive large-scale biomedical knowledge graph for AI-powered data-driven biomedical research | iKraph | Multi-source KG | Drug repurposing and Hypothesis Generation | Semi-automated | Nature Machine Intelligence | Link |
| 2025 | VITAGRAPH: Building a Knowledge Graph for Biologically Relevant Learning Tasks | VITAGRAPH | public KG | Drug repurposing | Semi-automated | arXiv | Link |
| 2024 | A Foundation Model for Clinician-Centered Drug Repurposing | / | public KG | Drug repurposing | Semi-automated | Nature Medicine | Link |
| 2024 | Accurate and Interpretable Drug-Drug Interaction Prediction Enabled by Knowledge Subgraph Learning | / | public KG | DDI prediction | Automated | Nature Communication Medicine | Link |
| 2024 | Knowledge Enhanced Representation Learning for Drug discovery | MKG | Multi-source KG | DTI prediction and Virtual screening and drug discovery | Semi-automated | AAAI | Link |
| 2024 | An experimentally validated approach to automated biological evidence generation in drug discovery using knowledge graphs | Healx KG | public KG | Drug repurposing | Semi-automated | Nature Communications | Link |
| 2024 | DDI-GPT: Explainable Prediction of Drug-Drug Interactions using Large Language Models enhanced with Knowledge Graphs | iBKH | public KG | DDI prediction | Semi-automated | bioRxiv | Link |
| 2024 | MKG-FENN: A Multimodal Knowledge Graph Fused End-to-End Neural Network for Accurate Drug–Drug Interaction Prediction | MKG | Multi-source KG | DDI prediction | Automated | AAAI | Link |
| 2024 | TransFOL: A Logical Query Model for Complex Relational Reasoning in Drug-Drug Interaction | / | public KG | DDI prediction | Semi-automated | Journal of Biomedical and Health Informatics | Link |
| 2024 | KGRLFF: Detecting Drug-Drug Interactions Based on Knowledge Graph Representation Learning and Feature Fusion | / | public KG | DDI prediction | Semi-automated | TCBB | Link |
| 2024 | An effective framework for predicting drug–drug interactions based on molecular substructures and knowledge graph neural network | DKG (Drug knowledge graph) | public KG | DDI prediction | Semi-automated | Computers in Biology and Medicine | Link |
| 2024 | Medical knowledge graph question answering for drug‐drug interaction prediction based on multi‐hop machine reading comprehension | / | public KG | DDI prediction | Automated | CAAI Transactions on Intelligence Technology | Link |
| 2024 | Integrated Knowledge Graph and Drug Molecular Graph Fusion via Adversarial Networks for Drug–Drug Interaction Prediction | DrugBank | public KG | DDI prediction | Semi-automated | JCIM | Link |
| 2024 | KGE-UNIT: toward the unification of molecular interactions prediction based on knowledge graph and multi-task learning on drug discovery | / | Multi-source KG | DDI prediction, DTI prediction and Hypothesis Generation | Automated | Briefings in Bioinformatics | Link |
| 2023 | Biomedical Knowledge Graph Learning for Drug Repurposing by Extending Guilt-By Association to Multiple Layers | / | public KG | Drug repurposing | Semi-automated | Nature Communications | Link |
| 2023 | Evolution-strengthened knowledge graph enables predicting the targetability and druggability of genes | ESKG (Evolution-strengthened KG) | public KG | DTI prediction | Semi-automated | PNAS nexus | Link |
| 2023 | Drugomics: Knowledge Graph & AI to Construct Physicians' Brain Digital Twin to Prevent Drug Side-Effects and Patient Harm | Drugomics KG | Multi-source KG | Drug toxicity and adverse reactions | Semi-automated | Big Data Analytics | Link |
| 2023 | Molecular-evaluated and explainable drug repurposing for COVID-19 using ensemble knowledge graph embedding | / | Multi-source KG | Drug repurposing | Semi-automated | Scientific Reports | Link |
| 2023 | Toxicology knowledge graph for structural birth defects | ReproTox-KG | Multi-source KG | Drug toxicity and adverse reactions and Hypothesis Generation | Semi-automated | Communications medicine | Link |
| 2023 | NAFLDkb: A Knowledge Base and Platform for Drug Development against Nonalcoholic Fatty Liver Disease | NAFLDkb | Multi-source KG | Drug repurposing | Semi-automated | JCIM | Link |
| 2023 | Molecule generation toward target protein (SARS-CoV-2) using reinforcement learning-based graph neural network via knowledge graph | / | domain-specific KG | Virtual screening and drug discovery, DTI prediction, and Hypothesis Generation | Semi-automated | Network Modeling and Analysis in Health Informatics and Bioinformatics | Link |
| 2022 | e-TSN: an Interactive Visual Exploration Platform for Target-Disease Knowledge Mappling from Literature | e-TSN KG | literature-based KG | DTI prediction | Automated | Briefings in Bioinformatics | Link |
| 2022 | Attention-based knowledge graph representation learning for predicting drug-drug interactions | / | public KG | DDI prediction | Semi-automated | Briefings in Bioinformatics | Link |
| 2022 | Automating Predictive Toxicology Using ComptoxAI | ComptoxAI KG | public KG | Drug toxicity and adverse reactions | Semi-automated | Chemical Research in Toxicology | Link |
| 2022 | KG-MTL: Knowledge Graph Enhanced Multi-Task Learning for Molecular Interaction | DRKG | public KG | DTI prediction | Automated | IEEE | Link |
| 2021 | A Unified Drug-Target Interaction Prediction Framework Based on Knowledge Graph and Recommendation System | / | public KG | DTI prediction | Semi-automated | Nature Communications | Link |
| 2021 | Biological Insights Knowledge Graph: an integrated knowledge graph to support drug development | BIKG | Multi-source KG | DTI prediction and Drug repurposing | Semi-automated | bioRxiv | Link |
| 2021 | SumGNN: multi-typed drug interaction prediction via efficient knowledge graph summarization | / | public KG | DDI prediction | Semi-automated | Bioinformatics | Link |
| 2021 | Predicting Potential Drug Targets Using Tensor Factorisation and Knowledge Graph Embeddings | Hetionet | public KG | DTI prediction | Semi-automated | BIOKDD | Link |
| 2021 | Adverse Drug Reaction Discovery Using a Tumor-Biomarker Knowledge Graph | TBKG (Tumor-biomarker KG) | literature-based KG | Drug toxicity and adverse reactions | Semi-automated | Frontiers in Genetics | Link |
| 2021 | Investigating ADR mechanisms with Explainable AI: a feasibility study with knowledge graph mining | PGxLOD | Multi-source KG | Drug toxicity and adverse reactions | Semi-automated | BMC Medical Informatics and Decision Making | Link |
| 2020 | Discovering Protein Drug Targets Using Knowledge Graph Embeddings | a knowledge graph of biological entities related to both drugs and targets | public KG | DTI prediction | Automated | Bioinformatics | Link |
| 2020 | KGNN: Knowledge Graph Neural Network for Drug-Drug Interaction Prediction | / | public KG | DDI prediction | Semi-automated | IJCAI | Link |
| 2020 | Network-based prediction of drug–target interactions using an arbitrary-order proximity embedded deep forest | / | public KG | DTI prediction | Semi-automated | Bioinformatics | Link |
| 2019 | Evaluation of knowledge graph embedding approaches for drug-drug interaction prediction in realistic settings | / | public KG | DDI prediction | Semi-automated | BMC Bioinformatics | Link |
| 2019 | Drug-Drug Interaction Prediction Based on Knowledge Graph Embeddings and Convolutional-LSTM Network | / | public KG | DDI prediction | Semi-automated | arXiv | Link |
| 2019 | GAMENet: Graph Augmented MEmory Networks for Recommending Medication Combination | EHR&DDI Graph | domain-specific KG | Virtual screening and drug discovery | Automated | AAAI | Link |
| 2019 | Facilitating prediction of adverse drug reactions by using knowledge graphs and multi‐label learning models | Bio2RDF KG | public KG | Drug toxicity and adverse reactions | Automated | Briefings in Bioinformatics | Link |
| 2018 | Modeling polypharmacy side effects with graph convolutional networks | / | public KG | Drug toxicity and adverse reactions | Semi-automated | Bioinformatics | Link |
| 2018 | Neural networks for link prediction in realistic biomedical graphs: a multi-dimensional evaluation of graph embedding-based approaches | / | public KG | DTI prediction | Semi-automated | BMC Bioinformatics | Link |
| 2017 | A Network Integration Approach for Drug-Target Interaction Prediction and Computational Drug Repositioning from Heterogeneous Information | / | public KG | DTI prediction and Drug repurposing | Semi-automated | Nature Communications | Link |
| 2017 | Knowledge graph prediction of unknown adverse drug reactions and validation in electronic health records | / | public KG | Drug toxicity and adverse reactions | Semi-automated | Scientific Reports | Link |
| 2017 | Large-scale structural and textual similarity-based mining of knowledge graph to predict drug–drug interactions | / | public KG | DDI prediction | Semi-automated | Journal of Web Semantics | Link |
| 2017 | Deep mining heterogeneous networks of biomedical linked data to predict novel drug–target associations | LTN (Linked Tripartite Network) | public KG | DTI prediction | Semi-automated | Bioinformatics | Link |
Omics Interpretation and Analysis
| Year | Title | KG Name | KG Type | Domain | Construction Method | Venue | Paper |
|---|---|---|---|---|---|---|---|
| 2025 | A novel approach for target deconvolution from phenotype-based screening using knowledge graph | P53_HUMAN PPIKG | public KG | Proteomics research | Semi-automated | Scientific Reports | Link |
| 2025 | Unified Knowledge-Guided Molecular Graph Encoder with multimodal fusion and multi-task learning | Elemental KG and Biological KG | Multi-source KG | Proteomics research | Semi-automated | Neural Networks | Link |
| 2025 | PhenoKG: Knowledge Graph-Driven Gene Discovery and Patient Insights from Phenotypes Alone | PhenoKG | public KG | Genomics research | Semi-automated | arXiv | Link |
| 2024 | Petagraph: A large-scale unifying knowledge graph framework for integrating biomolecular and biomedical data | Petagraph | Multi-source KG | Genomics research | Semi-automated | Scientific Data | Link |
| 2024 | An ontology-based knowledge graph for representing interactions involving RNA molecules | RNA-KG | Multi-source KG | Transcriptomics research | Semi-automated | Scientific Data | Link |
| 2024 | Knowledge graph construction based on granulosa cells transcriptome from polycystic ovary syndrome with normoandrogen and hyperandrogen | causal KG | Multi-source KG | Transcriptomics research | Semi-automated | Journal of Ovarian Research | Link |
| 2024 | Multi-Modal Protein Knowledge Graph Construction and Applications (Student Abstract) | ProteinKG65 | Multi-source KG | Proteomics research | Semi-automated | AAAI | Link |
| 2024 | Bridging chemical structure and conceptual knowledge enables accurate prediction of compound-protein interaction | DRKG | public KG | Proteomics research | Manual | BMC Biology | Link |
| 2024 | Integration of chromosome locations and functional aspects of enhancers and topologically associating domains in knowledge graphs enables versatile queries about gene regulation | crm, crm2gene, crm2tfac, crm2phen, tad, human genes (after ampliation) graph | public KG | Genomics research | Semi-automated | Nucleic Acids Research | Link |
| 2024 | Identifying compound-protein interactions with knowledge graph embedding of perturbation transcriptomics | / | public KG | Proteomics research | Semi-automated | Cell Genomics | Link |
| 2023 | MMiKG: a knowledge graph-based platform for path mining of microbiota–mental diseases interactions | MMiKG | literature-based KG | Microbiome research | Manual | Briefings in Bioinformatics | Link |
| 2023 | Transporter proteins knowledge graph construction and its application in drug development | Transporter Proteins Knowledge Graph | public KG | Proteomics research | Semi-automated | Computational and Structural Biotechnology Journal | Link |
| 2023 | A Knowledge Graph Approach to Elucidate the Role of Organellar Pathways in Disease via Biomedical Reports | / | Multi-source KG | Proteomics research | Semi-automated | JoVE Journal of Biochemistry | Link |
| 2022 | Knowledge-graph-based cell-cell communication inference for spatially resolved transcriptomic data with SpaTalk | LRT-KG | public KG | Transcriptomics research | Semi-automated | Nature Communications | Link |
| 2022 | A knowledge graph to interpret clinical proteomics data | CKG (Clinical Knowledge graph) | Multi-source KG | Proteomics research | Semi-automated | Nature Biotechnology | Link |
| 2022 | Knowledge integration and decision support for accelerated discovery of antibiotic resistance genes | E. coli knowledge graph | public KG | Genomics research | Semi-automated | Nature Communications | Link |
| 2022 | Machine learning prediction and tau-based screening identifies potential Alzheimer's disease genes relevant to immunity | PKG (protein knowledge graph) | public KG | Proteomics research | Semi-automated | Communications Biology | Link |
| 2022 | OntoProtein: Protein Pretraining With Gene Ontology Embedding | ProteinKG25 | public KG | Proteomics research | Automated | ICLR | Link |
| 2022 | GenomicKB: a knowledge graph for the human genome | GenomicKB (Genomic Knowledgebase) | Multi-source KG | Genomics research | Semi-automated | Nucleic Acids Research | Link |
| 2022 | Creating and Exploiting the Intrinsically Disordered Protein Knowledge Graph (IDP-KG) | IDP-KG | public KG | Proteomics research | Automated | CEUR Workshop Proceedings | Link |
| 2022 | BioTAGME: A Comprehensive Platform for Biological Knowledge Network Analysis | BioTAGME KG | Multi-source KG | Multi-Omics research | Semi-automated | Frontiers in Genetics | Link |
| 2022 | Biomedical knowledge graph embeddings for personalized medicine: Predicting disease-gene associations | a biomedical KG for predicting disease-gene association | public KG | Genomics research | Semi-automated | Expert Systems | Link |
| 2022 | Identifying genes targeted by disease-associated non-coding SNPs with a protein knowledge graph | Protein KG | literature-based KG | Genomics research | Semi-automated | PLOS ONE | Link |
| 2022 | KG-MTL: Knowledge Graph Enhanced Multi-Task Learning for Molecular Interaction | DRKG | public KG | Proteomics research | Automated | IEEE | Link |
| 2021 | FORUM: building a Knowledge Graph from public databases and scientific literature to extract associations between chemicals and diseases | FORUM | Multi-source KG | Metabolomics research | Semi-automated | Bioinformatics | Link |
| 2020 | Exploring the Microbiota-Gut-Brain Axis for Mental Disorders with Knowledge Graphs | MiKG (Microbiota knowledge graph) | Multi-source KG | Microbiome research | Semi-automated | Journal of Artificial Intelligence for Medical Sciences | Link |
| 2020 | Metastatic Site Prediction in Breast Cancer using Omics Knowledge Graph and Pattern Mining with Kirchhoff's Law Traversal | Kirchhoff's KG | public KG | Multi-Omics research | Semi-automated | bioRxiv | Link |
| 2020 | Accurate prediction of kinase-substrate networks using knowledge graphs | a phosphorylation knowledge graph | public KG | Proteomics research | Automated | PLoS Computational Biology | Link |
| 2020 | An integrative knowledge graph for rare diseases, derived from the Genetic and Rare Diseases Information Center (GARD) | an integrative KG for rare diseases | Multi-source KG | Genomics research | Semi-automated | Journal of Biomedical Semantics | Link |
| 2019 | Predicting gene-disease associations from the heterogeneous network using graph embedding | / | public KG | Genomics research | Semi-automated | IEEE | Link |
| 2019 | GenomicsKG: A Knowledge Graph to Visualize Poly-Omics Data | GenomicsKG | public KG | Genomics research | Semi-automated | Journal of Advances in Health | Link |
| 2018 | Semantic Disease Gene Embeddings (SmuDGE): phenotype-based disease gene prioritization without phenotypes | / | public KG | Genomics research | Semi-automated | Bioinformatics | Link |
| 2018 | Heterogeneous network embedding for identifying symptom candidate genes | SDGNet&SDGPNet (two heterogeneous symptom-related networks) | public KG | Genomics research | Semi-automated | JAMIA | Link |
| 2018 | Network-based integration of multi-omics data for prioritizing cancer genes | / | public KG | Genomics research | Semi-automated | Bioinformatics | Link |
| 2016 | A knowledge-based approach for predicting gene–disease associations | / | public KG | Genomics research | Semi-automated | Bioinformatics | Link |
Chemical Reaction and Synthesis
| Year | Title | KG Name | KG Type | Domain | Construction Method | Venue | Paper |
|---|---|---|---|---|---|---|---|
| 2025 | Bi-level Contrastive Learning for Knowledge-Enhanced Molecule Representations | MolKG | public KG | Molecular property prediction | Semi-automated | AAAI | Link |
| 2025 | Automated Retrosynthesis Planning of Macromolecules Using Large Language Models and Knowledge Graphs | / | literature-based KG | Chemical synthesis pathway optimization | Automated | Macromolecular Rapid Communications | Link |
| 2025 | An Automated Approach for Domain-Specific Knowledge Graph Generation─Graph Measures and Characterization | / | literature-based KG | Chemical reaction prediction and Chemical Synthesis Pathway Optimization | Automated | JCIM | Link |
| 2024 | Large-Scale Knowledge Integration for Enhanced Molecular Property Prediction | ElementKG-CHEBI | Multi-source KG | Molecular property prediction | Semi-automated | Nesy | Link |
| 2024 | Self-Supervised Contrastive Molecular Representation Learning with a Chemical Synthesis Knowledge Graph | Chemical synthesis KG | public KG | Chemical reaction prediction | Semi-automated | JCIM | Link |
| 2023 | Knowledge graph-enhanced molecular contrastive learning with functional prompt | ElementKG | domain-specific KG | Molecular property prediction | Semi-automated | Nature Machine Intelligence | Link |
| 2023 | Marie and BERT─A Knowledge Graph Embedding Based Question Answering System for Chemistry | TWA KG (the World Avatar KG) and Wikidata chemistry KG | Dynamic KG | Chemical reaction prediction and Molecular property prediction | Semi-automated | ACS Omega | Link |
| 2022 | MKGE: Knowledge graph embedding with molecular structure information | KCCR and DeepDDI | public KG | Molecular property prediction | Semi-automated | Computational Biology and Chemistry | Link |
| 2022 | Prediction of Compound Synthesis Accessibility Based on Reaction Knowledge Graph | Reaction knowledge graph | public KG | Chemical reaction prediction, Molecular property prediction, and Chemical synthesis pathway optimization | Semi-automated | Molecules | Link |
| 2022 | Molecular Contrastive Learning with Chemical Element Knowledge Graph | Chemical element KG | domain-specific KG | Molecular property prediction | Semi-automated | AAAI | Link |
| 2022 | FAIR and Interactive Data Graphics from a Scientific Knowledge Graph | / | literature-based KG | Chemical synthesis pathway optimization, Chemical reaction prediction, and Molecular property prediction | Semi-automated | Scientific Data | Link |
| 2022 | From Platform to Knowledge Graph: Evolution of Laboratory Automation | The World Avatar KG | Dynamic KG | Chemical Synthesis Pathway Optimization | Semi-automated | JACS Au | Link |
| 2021 | Intelligent generation of optimal synthetic pathways based on knowledge graph inference and retrosynthetic predictions using reaction big data | Reaction knowledge graph | public KG | Chemical synthesis pathway optimization | Semi-automated | Journal of the Taiwan Institute of Chemical Engineers | Link |
| 2021 | Automated Calibration of a Poly(oxymethylene) Dimethyl Ether Oxidation Mechanism Using the Knowledge Graph Technology | JPS KG | Dynamic KG | Chemical reaction prediction and Chemical Synthesis Pathway Optimization | Semi-automated | JCIM | Link |
| 2021 | A graph-based network for predicting chemical reaction pathways in solid-state materials synthesis | / | domain-specific KG | Chemical reaction prediction | Automated | Nature Communications | Link |
| 2020 | Knowledge Graph Approach to Combustion Chemistry and Interoperability | JPS KG | literature-based KG | Chemical reaction prediction | Automated | ACS Omega | Link |
| 2020 | Multiscale Cross-Domain Thermochemical Knowledge-Graph | JPS KG | Dynamic KG | Chemical reaction prediction and Molecular property prediction | Automated | JCIM | Link |
| 2016 | Modelling Chemical Reasoning to Predict and Invent Reactions | / | public KG | Chemical reaction prediction | Semi-automated | Chemistry Europe | Link |
Materials Design and Discovery
| Year | Title | KG Name | KG Type | Domain | Construction Method | Venue | Paper |
|---|---|---|---|---|---|---|---|
| 2025 | Construction of a knowledge graph for framework material enabled by large language models and its application | KG-FM | literature-based KG | Material screening and optimization | Semi-automated | npj Computational Materials | Link |
| 2025 | High throughput screening of new piezoelectric materials using graph machine learning and knowledge graph approach | a simple KG encoding structural similarity between materials | public KG | Material screening and optimization | Semi-automated | Computational Materials Science | Link |
| 2024 | MatKG: An autonomously generated knowledge graph in Material Science | MatKG | literature-based KG | New material design | Semi-automated | Scientific Data | Link |
| 2024 | A materials terminology knowledge graph automatically constructed from text corpus | MGED-KG | literature-based KG | Material screening and optimization | Semi-automated | Scientific Data | Link |
| 2024 | Construction and Application of Materials Knowledge Graph in Multidisciplinary Materials Science via Large Language Model | MKG | literature-based KG and dynamic KG | New material design | Semi-automated | NeurIPS | Link |
| 2024 | Generative Retrieval-Augmented Ontologic Graph and Multiagent Strategies for Interpretive Large Language Model-Based Materials Design | Ontological KG | literature-based KG | New material design and Material performance prediction | Semi-automated | ACS Engineering Au | Link |
| 2024 | SciAgents: Automating Scientific Discovery Through Bioinspired Multi-Agent Intelligent Graph Reasoning | an ontological knowledge graph for biologically inspired materials | literature-based KG | New material design and Material performance prediction | Semi-automated | Advanced Materials | Link |
| 2024 | An ontology-based text mining dataset for extraction of process-structure-property entities | Materials mechanics ontology | literature-based KG | Material performance prediction | Semi-automated | Scientific Data | Link |
| 2024 | Material Property Prediction with Element Attribute Knowledge Graphs and Multimodal Representation Learning | Element KG | domain-specific KG | Material performance prediction | Semi-automated | arXiv | Link |
| 2024 | Knowledge graph-guided data-driven design of ultra-high-performance concrete (UHPC) with interpretability and physicochemical reaction discovery capability | UHPC KG | literature-based KG | Material screening and optimization | Manual | Construction and Building Materials | Link |
| 2023 | The materials experiment knowledge graph | MekG | domain-specific KG | Material performance prediction | Semi-automated | Digital Discovery | Link |
| 2023 | Revisiting Electrocatalyst Design by a Knowledge Graph of Cu-Based Catalysts for CO2 Reduction | Cu-Based Catalysts Knowledge Graph for CO2 Reduction | literature-based KG | New material design and Material performance prediction | Semi-automated | ACS Catalysis | Link |
| 2023 | Reinforcement learning-based knowledge graph reasoning for aluminum alloy applications | Aluminum alloy domain KG | public KG | Material performance prediction | Semi-automated | Computational Materials Science | Link |
| 2023 | Bridging the Semantic-Numerical Gap: A Numerical Reasoning Method of Cross-modal Knowledge Graph for Material Property Prediction | Cross-modal KG | Multi-source KG | Material performance prediction | Semi-automated | arXiv | Link |
| 2023 | Digital Twin-Based Fault Diagnosis Platform for Final Rolling Temperature in Hot Strip Production | / | domain-specific KG | Material screening and optimization | Semi-automated | Materials | Link |
| 2022 | Grain Knowledge Graph Representation Learning: A New Paradigm for Microstructure-Property Prediction | Grain KG | domain-specific KG | Material performance prediction | Semi-automated | Crystals | Link |
| 2022 | Automating Materials Exploration with a Semantic Knowledge Graph for Li-Ion Battery Cathodes | a semantic knowledge graph dedicated to LIB cathodes | literature-based KG | New material design | Semi-automated | AFM | Link |
| 2022 | High-Throughput Computing Assisted by Knowledge Graph to Study the Correlation between Microstructure and Mechanical Properties of 6XXX Aluminum Alloy | 6XXX Aluminum Alloy KG | domain-specific KG | Material screening and optimization and Material performance prediction | Semi-automated | Materials | Link |
| 2022 | FAIR and Interactive Data Graphics from a Scientific Knowledge Graph | / | literature-based KG | Material screening and optimization | Semi-automated | Scientific Data | Link |
| 2022 | Compound Knowledge Graph-Enabled AI Assistant for Accelerated Materials Discovery | CKG (Compound KG) | Multi-source KG | New material design | Semi-automated | Integrating Materials and Manufacturing Innovation | Link |
| 2021 | EBSD Grain Knowledge Graph Representation Learning for Material Structure-Property Prediction | EBSD Grain KG | domain-specific KG | Material performance prediction | Semi-automated | CCKS | Link |
| 2020 | propnet: A Knowledge Graph for Materials Science | propnet KG | domain-specific KG | Material performance prediction | Semi-automated | Matter | Link |
| 2020 | NanoMine: A Knowledge Graph for Nanocomposite Materials Science | NanoMine KG | domain-specific KG | New material design | Semi-automated | ISWC | Link |
| 2018 | Relation extraction with weakly supervised learning based on process-structure-property-performance reciprocity | PSPP KG (Process-Structure-Property-Performance) | literature-based KG | New material design | Semi-automated | Science and Technology of Advanced Materials | Link |
