Ontology
Introduction
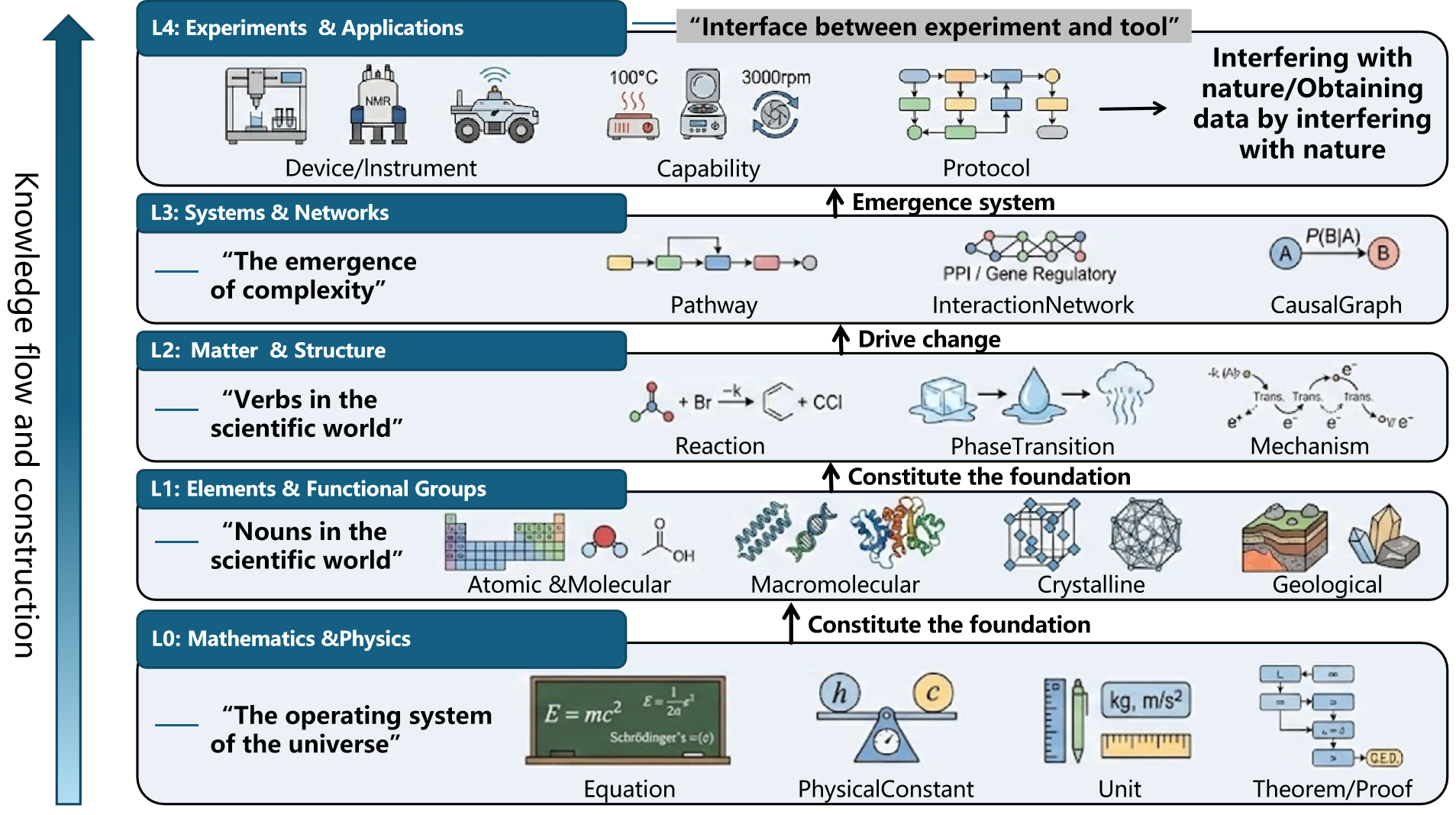
To bridge the gap between the probabilistic intuition of LLMs and the deterministic logic of the physical world, we propose a unified representation framework for scientific knowledge. Instead of using disciplinary boundaries (physics/chemistry/biology) as a means of knowledge organization, this framework adopts an orthogonal hierarchical architecture based on abstraction levels, decomposing scientific knowledge into five mutually independent and vertically coupled levels: Layer 0 (L0) - mathematical and physical foundation layer, Layer 1 (L1) - static material ontology layer, Layer 2 (L2) - dynamic mechanisms and processes layer, Layer 3 (L3) - networks and systems layer, and Layer 4 (L4) - execution and capability layer.
SciOntology
✨ SciOntology (Unified Framework for Scientific Knowledge Representation) breaks disciplinary silos (physics, chemistry, biology, materials) and introduces an orthogonal layered architecture that elevates scientific knowledge from an encyclopedia-style listing to a computational cognitive operating system. The framework consists of five vertically coupled yet independent layers: L0 Mathematical & Physical Foundations → L1 Static Matter Ontology → L2 Dynamics & Mechanisms → L3 Networks & Systems → L4 Execution & Capabilities. This structure guarantees physical plausibility, structural legality, and causal coherence in scientific reasoning.
Provides a formal, computable symbol system: equations, physical constants, units/dimensions, theorems, and proofs. Using the Physics Derivation Graph (PDG) and autoformalization (Lean/Coq), this layer projects LLM outputs from “linguistically plausible” to “physically admissible”. It enforces conservation laws, dimensional consistency, and symbolic derivation validity, eliminating numerical or unit hallucinations.
All independently existing entities: atoms/molecules, macromolecules (proteins, nucleic acids), crystalline materials, geological features, and more. Following the identity stability principle, each entity retains a unique node regardless of context. Unified through ChEBI, PRO, CSO, and other ontologies, L1 provides precise symbolic anchors for entity linking and cross‑domain transfer (e.g., graphene's conductivity from materials to biosensors).
Explicitly distinguishes reactions (input/output mapping), phase transitions (physical state change), and mechanisms (explanatory layer). Tracks electron redistribution (arrow pushing), transition states, and energy barriers. As occurrents with temporal parts, L2 supports time-series modeling and experimental sequence logic (e.g., “add acid before water” vs. reverse). Enables AI to understand how changes happen, not just what changes.
Models pathways, protein-protein interaction networks, and causal graphs to capture emergent behaviors that cannot be reduced to individual components. Introduces CausalRAG — retrieval of causal paths rather than text chunks — supporting counterfactual reasoning and explanatory chains. This shifts AI from associative correlation to causal inference, enabling discovery of non-linear “butterfly effect” mechanisms in systems biology, Earth system science, and complex materials.
Abstracts hardware into capabilities rather than specific devices. Based on SiLA 2 (microservice communication) and LabOP (semantic protocol standards), L4 translates high-level scientific intent into hardware-agnostic executable protocols. The AI agent closes the loop: plan generation → capability matching → instruction compilation (SiLA 2) → sensor feedback (SSN/SOSA) → state update. Embeds safety constraints as hard boundaries for autonomous experimentation.
