SCP-KG
@ZJU
ElementKG is a knowledge graph centered on chemical elements, designed to summarize fundamental chemical knowledge and cover elements as well as their closely related functional groups.
Chemistry

@ZJU
ElementKG 2.0 is a chemistry knowledge graph that spans the entire chain from “elements–functional groups–molecules–reactions–experiments,” aiming to model the full lifecycle of chemical reactions.
Chemistry
@ZJU
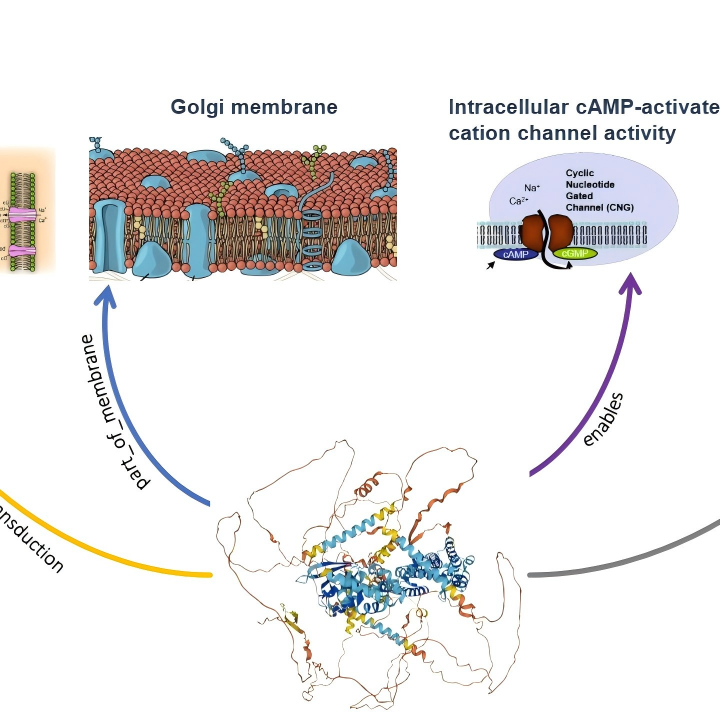
ProteinKG65 is a large-scale knowledge graph dataset for protein science. It aligns protein sequences with functional and ontological knowledge (from Gene Ontology, GO) and is compatible with structured knowledge, sequences, and textual descriptions.
Biology
@ZJU
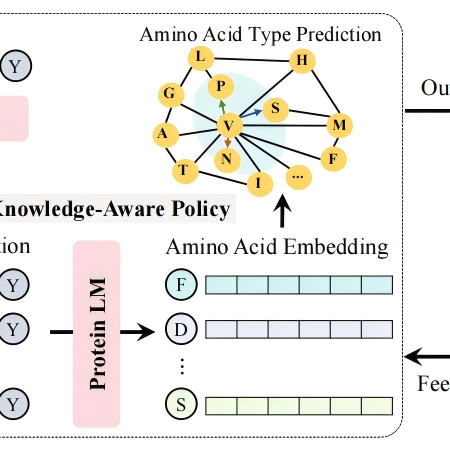
Amino Acid Knowledge Graph (AAKG) is a domain-specific knowledge graph that systematically represents the biochemical properties of amino acids and their intrinsic relationships.
Biology

@ZJU
PorousKG is a knowledge graph of porous materials centered on metal–organic frameworks (MOFs) and extended to covalent organic frameworks (COFs), hydrogen-bonded organic frameworks (HOFs), and cage-like materials.
Materials

@ZJU
MolSafeKG is a heterogeneous knowledge graph designed for molecular safety assessment. It integrates 83,925 hazardous molecules from authoritative sources such as ECHA and ChEMBL,
Chemistry

@ZJU
OneGraph is a general-purpose knowledge graph constructed with the assistance of large language models. It spans common-sense concepts and relations across the humanities, engineering and technology,
Biology

@ZJU
TTG-Math is a structured knowledge graph designed for mathematical reasoning. It abstracts mathematical problems into general problem templates and solution templates,
Chemistry

@ZJU
InstructProteinKG is a protein knowledge graph tailored for protein sequence-text alignment and instruction learning. Primarily extracted from the high-quality
Biology

BEACON
@ZJU
BEACON (Bridging Chemical Structure and Conceptual Knowledge) is an innovative bioinformatics framework and its associated dataset, proposed in 2024 in BMC Biology. This project introduces a bidirectional fusion approach to address the bottleneck in compound-protein interaction (CPI) prediction, where traditional methods often view chemical structures or biological background knowledge in isolation.
Chemistry

DREAMwalk
@ZJU
The dataset utilized by DREAMwalk is a multi-layer heterogeneous biomedical knowledge graph. It was not constructed from scratch, but rather formed a foundational network by integrating multiple authoritative public databases (such as DrugBank, DisGeNET, STRING, etc.), and dynamically incorporated semantic information on this basis. Its core innovation lies in utilizing the hierarchical structure of Ontology to calculate semantic similarities among entities of the same type, and incorporating these similarities as new “virtual edges” into the original graph, thereby creating a richer and more biologically semantic-capturing enhanced knowledge graph specifically designed to improve the predictive performance of tasks such as drug repositioning.
Biology

HealxKG
@ZJU
HealxKG is a multi-source integrated biomedical knowledge graph for rare disease drug repositioning, developed by Healx Ltd. It aims to generate interpretable therapeutic hypotheses through symbolic reasoning and automated filtering mechanisms. The graph integrates public, commercial, and internally curated data, covering core entities such as compounds, diseases, genes, pathways, and their pharmacological, genetic, and clinical relationships. It has been experimentally validated in fragile X syndrome, cystic fibrosis, and Parkinson’s disease.
Chemistry

Material-KG
@ZJU
Material-Knowledge-Graph is not a directly downloadable “finished” knowledge graph (such as an RDF file or a Neo4j database), but rather an open-source project and accompanying dataset for automatically constructing a knowledge graph of materials science. Its core objective is to utilize a large language model (LLM) to automatically extract structured knowledge from a vast amount of materials science literature, thereby building a knowledge graph encompassing elements such as material names, chemical formulas, properties, and synthesis methods.
Materials

E. coliKG
@ZJU
The KIDS E. coli Antibiotic Resistance Knowledge Graph is a systems biology knowledge graph jointly developed by the Tagkopoulos Laboratory at UC Davis, Zhejiang University, and Fudan University. This graph focuses on antibiotic resistance (AMR) research across the entire genome of Escherichia coli (E. coli), aiming to accelerate the discovery of resistance genes through knowledge integration and automated reasoning.
Biology

OPB
@ZJU
OPB is a reference ontology that applies classical physics and thermodynamics to the dynamics of biological systems. It is designed to encompass multiple structural scales (from atoms to organisms) and various physical fields (fluid dynamics, chemical kinetics, particle diffusion, etc.) encountered in the research and analysis of biological organisms.
Biology

ProteinKG25
@ZJU
ProteinKG25 is a large-scale biomedical knowledge graph (KG) oriented towards understanding protein function, developed by the ZJUNLP team at Zhejiang University. It aims to support the pre-training of knowledge-enhanced protein language models. This graph integrates structured ontological relationships from Gene Ontology (GO) with high-quality protein sequences and annotation information from Swiss-Prot, covering core entities such as gene ontology terms, proteins, and their semantic associations.
Biology

RNA-KG
@ZJU
RNA-KG v2.0 is an RNA-centered heterogeneous biomedical knowledge graph (KG). It not only integrates traditional “entity-relationship” information but also introduces contextual properties. It addresses the common issue of “disconnection from context” in biomedical data, enabling precise description of the specific biological contexts in which RNA interactions occur.
Biology

SSC-CoT-KG
@ZJU
The SSC-CoT-KG is a structured knowledge base specifically designed for mathematical reasoning in large language models (LLM). It focuses on formula derivation and logical path retrieval in the fields of trigonometry and geometry. It addresses the common “logical leap” problem encountered by large models in complex mathematical reasoning, where the model may be interrupted in the multi-step derivation process due to forgetting or inability to recall key formulas. The knowledge graph guides the model to construct a complete and coherent reasoning chain by providing precise formula clues for intermediate steps.
Mathematics

TxGNN KG
@ZJU
TxGNN is a large-scale biomedical knowledge graph (KG) specifically designed for predicting the relationship between drug repositioning and disease treatment. This graph integrates structured knowledge from multiple biomedical databases and literature, covering core biomedical entities such as drugs, diseases, genes, proteins, pathways, and their relationships. It has been successfully imported into the Neo4j graph database, containing approximately 129,312 nodes and 7,708,348 relationships.
Chemistry

TransFOL
@ZJU
TransFOL is a multi-source fused biomedical knowledge graph designed for predicting Drug-Drug Interactions (DDIs) and inferring complex relationships. This knowledge graph integrates structured knowledge from four authoritative data sources (DrugBank, TWOSIDES, DrugCombDB, Phenomebrowser), encompassing core biomedical entities such as drugs, proteins, enzymes, biological pathways, diseases, and phenotypes. It supports deep learning-driven DDI prediction and complex logical query reasoning, and holds significant application value in clinical drug safety assessment, drug repositioning, and precision medicine.
Chemistry

SpaTalk
@ZJU
SpaTalk is a knowledge graph-based dataset for inferring cell communication, developed by Zhejiang University, specifically designed for spatial transcriptomic data analysis. This graph constructs a three-tier knowledge framework of “ligand-receptor-target”, encompassing both human and mouse species, including 3,324 pairs of human and 2,484 pairs of mouse ligand-receptor pairs, as well as tens of thousands of intracellular signaling pathways. The dataset integrates multiple database resources such as CellTalkDB, KEGG, Reactome, and AnimalTFDB, systematically describing the complete transmission process of cell-to-cell communication signals from extracellular ligands to intracellular target genes through four relationship types: ligand-receptor binding, intracellular interaction, pathway assignment, and transcription factor annotation. This graph is primarily applied to inferring the communication intensity between adjacent cells in spatial transcriptomic data, tracing intracellular signaling pathways, and studying tissue microenvironment, supporting the data analysis of various spatial transcriptomic technologies such as STARmap, Slide-seq, and 10X Visium.
Biology

PhySci
@ZJU
PhySci is an ontology dedicated to scientific publications in physics research. By defining the overall structure of scientific publications, it constructs a knowledge base representation in the field of physics research. Utilizing the principles of Linked Data, it is developed with metadata concepts such as scientific methods, scientific problems, and solutions, achieving an interactive semantic model for heterogeneous data across sub-fields.
Biology

PertKGE
@ZJU
PertKGE is a compound-protein interaction prediction dataset released by the Shanghai Institute of Materia Medica, Chinese Academy of Sciences, and Tsinghua University in 2024. This atlas innovatively constructs a fine-grained knowledge graph that integrates perturbation transcriptomics. By dismantling genes into multiple levels of nucleic acid entities such as DNA, mRNA, lncRNA, and miRNA, and combining compounds, proteins, and transcription factors, it systematically describes the central dogma process of biological systems. The dataset integrates large-scale transcriptome perturbation data such as CMap/LINCS with various public biological network resources, containing approximately 80,000 entities and over 6.8 million triples, including 4.36 million regulatory relationships between compounds and mRNA, and 15,000 physical binding relationships between compounds and proteins. This atlas is primarily applied to new compound target prediction, ligand virtual screening, and drug action mechanism analysis. It effectively mitigates systematic biases in transcriptome data through knowledge graph embedding technology, enhancing the biological interpretability of predictions.
Biology

Cisreg
@ZJU
Cisreg is a gene regulatory knowledge graph dataset integrating chromosome location and function, released by institutions such as the University of Murcia in Spain and the Norwegian University of Science and Technology in 2024. The graph is built based on the Resource Description Framework standard, integrating 25 enhancer databases and 2 topological association domain resources, systematically describing the complex regulatory relationships between entities such as enhancers, topological association domains, genes, transcription factors, and phenotypes. The dataset comprises nearly 1 billion triples, including approximately 683 million pieces of enhancer sequence metadata, 263 million enhancer-target gene associations, and 3.8 million pieces of topological association domain structural information. The graph deeply integrates enhancer sequences, genomic coordinates, and biological functional semantics (such as transcription factor binding and phenotype association), supporting complex regulatory queries that combine spatial position constraints with functional semantics. It is primarily applied to the analysis of long-range gene expression regulatory mechanisms, the assessment of the impact of mutations in non-coding regions, and interoperability research across enhancer databases.
Biology

CKG
@ZJU
CKG is an open-source, large-scale biomedical knowledge graph platform that provides a complete database dump file, containing over 16 million nodes and 220 million relationships. This graph integrates nine biomedical ontologies, 26 public databases, and published experimental research data, unifying vast amounts of heterogeneous biomedical information into a graph structure. Its main application value lies in accelerating proteomics data analysis, supporting clinical decision-making (such as biomarker discovery), and facilitating knowledge discovery. It enables users to uncover hidden associations between entities such as genes, proteins, diseases, and drugs through graph database queries. The dataset is released in the form of a Neo4j database dump file, allowing users to set up a fully functional graph database instance locally for exploration.
Biology

IDP-KG
@ZJU
IDP-KG is an integrated knowledge graph dataset of intrinsically disordered proteins released by the University of Padova, Heriot-Watt University, and the ELIXIR IDP community in 2022. This graph adopts a fully automated ETL process, integrating the Bioschemas annotation data from three professional databases: DisProt, MobiDB, and PED. It unifies entities such as intrinsically disordered proteins, sequence annotations, sequence ranges, and IDPO ontological terms into the RDF standard format. The dataset contains 149 different IDPO codes, with 7,542 disorder annotations and 1,325 protein binding annotations, and fully records the source and traceability information of the data. This graph is primarily used to address the issue of inconsistent identifiers among disordered protein databases, supports cross-database federated queries and in-depth functional analysis, provides high-quality training labels for machine learning models, and serves as a benchmark dataset for the large-scale application of Bioschemas standards in the field of life sciences.
Biology

KG-FM
@ZJU
KG-FM is a large-scale framework material knowledge graph constructed by the MontageBai team between 2024 and 2025, specifically designed for metal-organic frameworks, covalent organic frameworks, and hydrogen-bonded organic frameworks. This graph utilizes a large language model to automatically extract structured knowledge from over 100,000 academic papers, constructing a knowledge network comprising 2.53 million nodes and 4.01 million relationships, covering core entity types such as framework materials, synthesis methods, performance indicators, application fields, chemical components, and literature. The graph supports multi-hop reasoning queries through semantic relationships such as has_synthesis_method, exhibits_performance, and used_for. It enables precise Cypher retrieval based on the Neo4j graph database and integrates an intelligent question-answering system based on the Qwen2 large model, providing answers with traceable literature citations. It is primarily applied to rapid retrieval of framework material properties and applications, research trend analysis, hotspot mining, as well as AI-driven material discovery and inverse design.
Materials

MatKG
@ZJU
MatKG is a large-scale, comprehensive Knowledge Graph (KG) for materials science, developed by a research team at the Massachusetts Institute of Technology (MIT). Utilizing advanced Natural Language Processing (NLP) technology, it automatically extracts entities and relationships from over 5 million scientific papers in the field of materials science, constructing a structured knowledge base. This graph aims to address the issue of highly fragmented knowledge in the field of materials science, which is difficult to integrate and utilize, providing a robust data foundation for applications such as material discovery, recommendation systems, and advanced analysis.
Materials

MKG-FENN
@ZJU
MKG-FENN (Multimodal Knowledge Graph Fused End-to-end Neural Network) is a multimodal knowledge graph fusion system specifically designed for accurately predicting Drug-Drug Interactions (DDIs). This knowledge graph is based on the paper with the same name published at AAAI 2024.
MKG-FENN has constructed a comprehensive biomedical knowledge graph by integrating structured and semi-structured knowledge from multiple authoritative data sources, including drug information, protein targets, biological pathways, molecular characteristics, and clinical interaction data. This graph supports deep learning-driven DDI prediction and holds significant application value in clinical drug safety assessment, precision medicine, and new drug discovery.
Chemistry

OM
@ZJU
OM is a comprehensive unit of measure ontology that provides standardized representations for units, dimensions, and related concepts. It supports scientific computation, data exchange, and semantic integration, and is particularly suitable for scenarios requiring precise unit conversion and dimensional analysis.
Physics

ADR-Graph
@ZJU
EdgePrediction is a machine learning library implemented in Python, specifically designed for Knowledge Graph Completion, particularly for Edge Prediction tasks. This library implements the algorithm described in Bean et al.’s (2017) paper, aiming to predict missing new members within a given category (i.e., predict new edges) by training a binary classifier. The algorithm was originally designed and applied for predicting Adverse Drug Reactions (ADRs).
The core logic involves utilizing Fisher’s exact test to identify the enriched attributes of positive examples in the training data as predictive features, and determining feature weights through grid search to maximize the objective function (default is Youden’s J statistic), thereby predicting the missing edges in the graph.
Chemistry

DDKG
@ZJU
DDKG (Attention-based Knowledge Graph Representation Learning for Predicting Drug-drug Interactions) is a biomedical knowledge graph dataset and algorithmic framework dedicated to predicting drug-drug interactions (DDIs). Proposed by institutions such as the Xinjiang Institute of Physical and Chemical Technology, this research aims to address the issue that existing computational methods fail to fully utilize drug attributes (such as SMILES sequences) and complex triples in biomedical knowledge graphs. By integrating attention mechanisms and graph neural networks, DDKG achieves end-to-end prediction of drug interaction probabilities.
Chemistry

DNINet
@ZJU
DTINet (Drug-Target Interaction Network) is a computational framework and dataset for predicting novel drug-target interactions (DTIs). This method integrates multi-source information from heterogeneous biological networks, learns low-dimensional feature vector representations for drug and protein nodes, and predicts new drug-target interactions based on a vector space projection scheme. This dataset and method have been applied in drug repositioning and computational drug discovery research.
Chemistry

Otter UBC
@ZJU
Otter UBC is a biomedical knowledge graph released by the Otter Research team in 2023, focusing on protein-drug interactions in the field of drug discovery. The graph integrates three major biomedical resources: UniProt, BindingDB, and ChEMBL, containing 573,227 manually annotated proteins, over 1.2 million drug molecules, and approximately 2.23 million non-overlapping protein-drug interaction triples, totaling over 6.2 million relationships. The graph provides multimodal attribute data, including protein sequences, drug SMILES expressions, pathway information, and catalytic activity, and establishes cross-database identifier alignment between UniProt, ChEMBL, and DrugBank. The dataset is published in RDF Turtle format, supporting flexible retrieval through SPARQL query language, and is primarily used for drug target identification, protein-drug relationship mining, and multimodal biomedical data analysis in precision medicine.
Chemistry

Otter STITCH
@ZJU
Otter STITCH is a high-confidence chemical-protein interaction knowledge graph released by the Otter Research team in 2023, focusing on scalable graph neural network training in the field of drug discovery. The graph is constructed by filtering interaction triples with confidence scores ≥0.9 from the STITCH database version 5.0, containing over 10.71 million relationships, covering 17,572 chemical molecules and 1,886,496 proteins. The graph provides multimodal attribute data, including SMILES expressions for chemical molecules, protein sequences, and quantitative confidence scores, and integrates computational predictions, cross-species knowledge transfer, and multiple database evidence. To overcome the computational bottleneck of large-scale knowledge graphs, the dataset is divided into five equally sized subgraphs (each with approximately 2.1 million triples), supporting a sequential training strategy for graph neural networks. It is primarily applied to scalable drug-protein interaction prediction and large-scale biomedical relationship mining.
Chemistry

Otter PrimeKG
@ZJU
Otter PrimeKG is a comprehensive biomedical knowledge graph for precision medicine, released by the Otter Research team in 2023. This knowledge graph integrates 20 biomedical resources and constructs a large-scale knowledge network encompassing 17,080 diseases, 29,786 genes/proteins, and 7,957 drugs, with a total of over 12.7 million relationship edges. The graph covers 13 multimodal data types, including protein sequences, drug SMILES expressions, pathway information, and clinical annotations, and provides over 640,000 protein-protein interactions, over 25,000 drug-protein interactions, and over 2.67 million drug-drug interactions. This dataset is published in RDF format and supports SPARQL queries. It is primarily used in drug repositioning research, disease mechanism analysis, and pathway analysis, providing comprehensive disease-gene-drug association evidence for precision medicine research.
Chemistry

Otter DUDe
@ZJU
Otter DUDe is a high-confidence drug-target interaction benchmark dataset released by the Otter Research team in 2023, specifically designed for model validation and performance evaluation in the field of drug discovery. This atlas is carefully curated from the DUDe database, retaining 40,216 experimentally validated positive interaction pairs, covering 22,886 active compounds and 102 protein targets. The dataset undergoes a rigorous preprocessing workflow: all negative samples are removed, and overlaps with the TDC DTI benchmark dataset are eliminated, ensuring that only experimentally confirmed positive interactions are retained. The atlas provides multimodal attribute data, including drug SMILES expressions, protein sequences, and quantitative affinity scores such as IC50, Ki, and Kd, and is standardized using UniProt accession numbers and ChEMBL/DrugBank IDs for identification. The dataset is released in CSV/TSV formats and is primarily used for fair performance evaluation and benchmark testing of drug-target prediction models, providing clean, data-leakage-free training and testing data for computational drug discovery research.
Chemistry
@ZJU
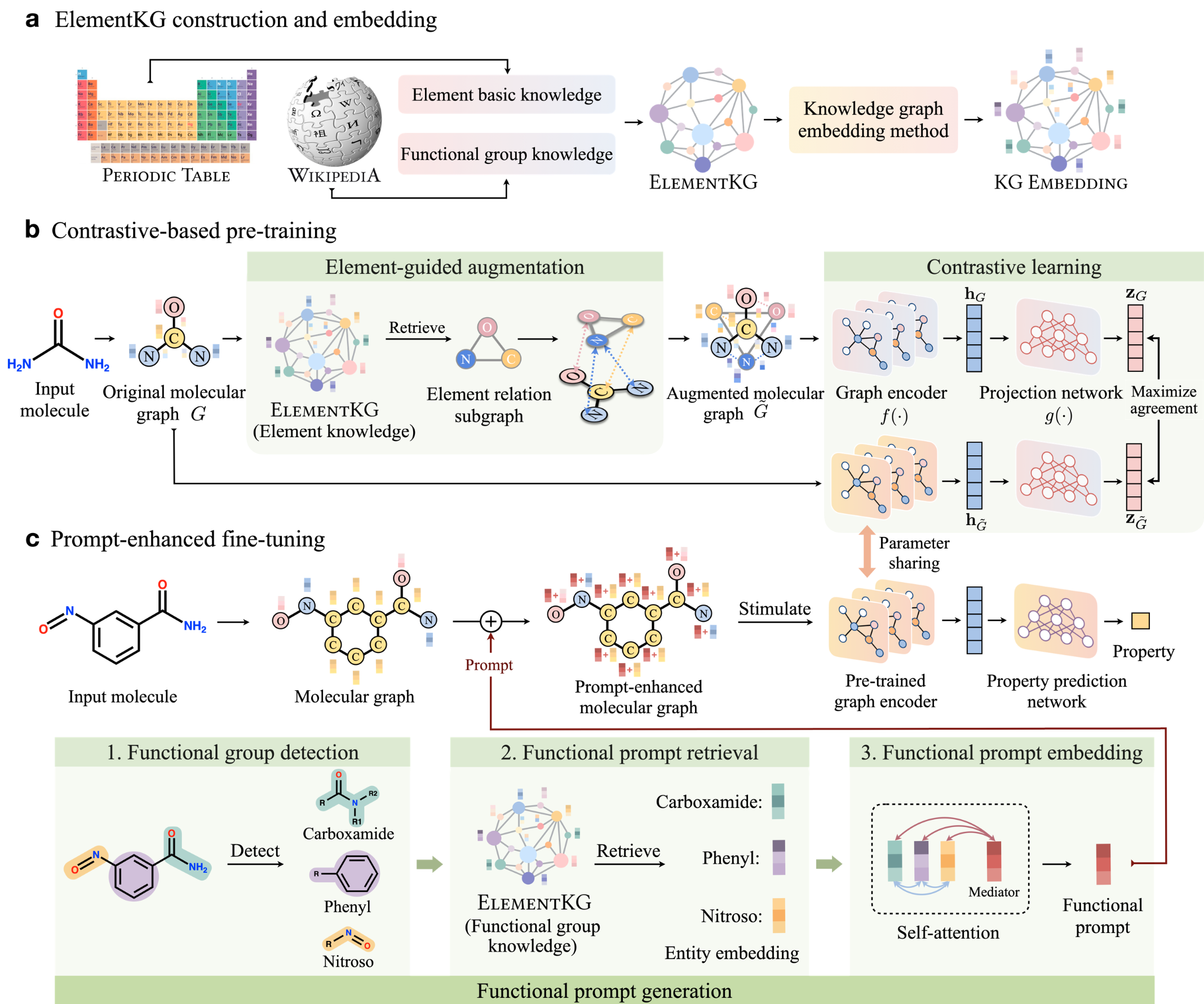
ElementKG is a knowledge graph centered on chemical elements, designed to summarize fundamental chemical knowledge and cover elements as well as their closely related functional groups.
Chemistry

@ZJU
ElementKG 2.0 is a chemistry knowledge graph that spans the entire chain from “elements–functional groups–molecules–reactions–experiments,” aiming to model the full lifecycle of chemical reactions.
Chemistry

@ZJU
MolSafeKG is a heterogeneous knowledge graph designed for molecular safety assessment. It integrates 83,925 hazardous molecules from authoritative sources such as ECHA and ChEMBL,
Chemistry

@ZJU
TTG-Math is a structured knowledge graph designed for mathematical reasoning. It abstracts mathematical problems into general problem templates and solution templates,
Chemistry

BEACON
@ZJU
BEACON (Bridging Chemical Structure and Conceptual Knowledge) is an innovative bioinformatics framework and its associated dataset, proposed in 2024 in BMC Biology. This project introduces a bidirectional fusion approach to address the bottleneck in compound-protein interaction (CPI) prediction, where traditional methods often view chemical structures or biological background knowledge in isolation.
Chemistry

HealxKG
@ZJU
HealxKG is a multi-source integrated biomedical knowledge graph for rare disease drug repositioning, developed by Healx Ltd. It aims to generate interpretable therapeutic hypotheses through symbolic reasoning and automated filtering mechanisms. The graph integrates public, commercial, and internally curated data, covering core entities such as compounds, diseases, genes, pathways, and their pharmacological, genetic, and clinical relationships. It has been experimentally validated in fragile X syndrome, cystic fibrosis, and Parkinson’s disease.
Chemistry

TxGNN KG
@ZJU
TxGNN is a large-scale biomedical knowledge graph (KG) specifically designed for predicting the relationship between drug repositioning and disease treatment. This graph integrates structured knowledge from multiple biomedical databases and literature, covering core biomedical entities such as drugs, diseases, genes, proteins, pathways, and their relationships. It has been successfully imported into the Neo4j graph database, containing approximately 129,312 nodes and 7,708,348 relationships.
Chemistry

TransFOL
@ZJU
TransFOL is a multi-source fused biomedical knowledge graph designed for predicting Drug-Drug Interactions (DDIs) and inferring complex relationships. This knowledge graph integrates structured knowledge from four authoritative data sources (DrugBank, TWOSIDES, DrugCombDB, Phenomebrowser), encompassing core biomedical entities such as drugs, proteins, enzymes, biological pathways, diseases, and phenotypes. It supports deep learning-driven DDI prediction and complex logical query reasoning, and holds significant application value in clinical drug safety assessment, drug repositioning, and precision medicine.
Chemistry

MKG-FENN
@ZJU
MKG-FENN (Multimodal Knowledge Graph Fused End-to-end Neural Network) is a multimodal knowledge graph fusion system specifically designed for accurately predicting Drug-Drug Interactions (DDIs). This knowledge graph is based on the paper with the same name published at AAAI 2024.
MKG-FENN has constructed a comprehensive biomedical knowledge graph by integrating structured and semi-structured knowledge from multiple authoritative data sources, including drug information, protein targets, biological pathways, molecular characteristics, and clinical interaction data. This graph supports deep learning-driven DDI prediction and holds significant application value in clinical drug safety assessment, precision medicine, and new drug discovery.
Chemistry

ADR-Graph
@ZJU
EdgePrediction is a machine learning library implemented in Python, specifically designed for Knowledge Graph Completion, particularly for Edge Prediction tasks. This library implements the algorithm described in Bean et al.’s (2017) paper, aiming to predict missing new members within a given category (i.e., predict new edges) by training a binary classifier. The algorithm was originally designed and applied for predicting Adverse Drug Reactions (ADRs).
The core logic involves utilizing Fisher’s exact test to identify the enriched attributes of positive examples in the training data as predictive features, and determining feature weights through grid search to maximize the objective function (default is Youden’s J statistic), thereby predicting the missing edges in the graph.
Chemistry

DDKG
@ZJU
DDKG (Attention-based Knowledge Graph Representation Learning for Predicting Drug-drug Interactions) is a biomedical knowledge graph dataset and algorithmic framework dedicated to predicting drug-drug interactions (DDIs). Proposed by institutions such as the Xinjiang Institute of Physical and Chemical Technology, this research aims to address the issue that existing computational methods fail to fully utilize drug attributes (such as SMILES sequences) and complex triples in biomedical knowledge graphs. By integrating attention mechanisms and graph neural networks, DDKG achieves end-to-end prediction of drug interaction probabilities.
Chemistry

DNINet
@ZJU
DTINet (Drug-Target Interaction Network) is a computational framework and dataset for predicting novel drug-target interactions (DTIs). This method integrates multi-source information from heterogeneous biological networks, learns low-dimensional feature vector representations for drug and protein nodes, and predicts new drug-target interactions based on a vector space projection scheme. This dataset and method have been applied in drug repositioning and computational drug discovery research.
Chemistry

Otter UBC
@ZJU
Otter UBC is a biomedical knowledge graph released by the Otter Research team in 2023, focusing on protein-drug interactions in the field of drug discovery. The graph integrates three major biomedical resources: UniProt, BindingDB, and ChEMBL, containing 573,227 manually annotated proteins, over 1.2 million drug molecules, and approximately 2.23 million non-overlapping protein-drug interaction triples, totaling over 6.2 million relationships. The graph provides multimodal attribute data, including protein sequences, drug SMILES expressions, pathway information, and catalytic activity, and establishes cross-database identifier alignment between UniProt, ChEMBL, and DrugBank. The dataset is published in RDF Turtle format, supporting flexible retrieval through SPARQL query language, and is primarily used for drug target identification, protein-drug relationship mining, and multimodal biomedical data analysis in precision medicine.
Chemistry

Otter STITCH
@ZJU
Otter STITCH is a high-confidence chemical-protein interaction knowledge graph released by the Otter Research team in 2023, focusing on scalable graph neural network training in the field of drug discovery. The graph is constructed by filtering interaction triples with confidence scores ≥0.9 from the STITCH database version 5.0, containing over 10.71 million relationships, covering 17,572 chemical molecules and 1,886,496 proteins. The graph provides multimodal attribute data, including SMILES expressions for chemical molecules, protein sequences, and quantitative confidence scores, and integrates computational predictions, cross-species knowledge transfer, and multiple database evidence. To overcome the computational bottleneck of large-scale knowledge graphs, the dataset is divided into five equally sized subgraphs (each with approximately 2.1 million triples), supporting a sequential training strategy for graph neural networks. It is primarily applied to scalable drug-protein interaction prediction and large-scale biomedical relationship mining.
Chemistry

Otter PrimeKG
@ZJU
Otter PrimeKG is a comprehensive biomedical knowledge graph for precision medicine, released by the Otter Research team in 2023. This knowledge graph integrates 20 biomedical resources and constructs a large-scale knowledge network encompassing 17,080 diseases, 29,786 genes/proteins, and 7,957 drugs, with a total of over 12.7 million relationship edges. The graph covers 13 multimodal data types, including protein sequences, drug SMILES expressions, pathway information, and clinical annotations, and provides over 640,000 protein-protein interactions, over 25,000 drug-protein interactions, and over 2.67 million drug-drug interactions. This dataset is published in RDF format and supports SPARQL queries. It is primarily used in drug repositioning research, disease mechanism analysis, and pathway analysis, providing comprehensive disease-gene-drug association evidence for precision medicine research.
Chemistry

Otter DUDe
@ZJU
Otter DUDe is a high-confidence drug-target interaction benchmark dataset released by the Otter Research team in 2023, specifically designed for model validation and performance evaluation in the field of drug discovery. This atlas is carefully curated from the DUDe database, retaining 40,216 experimentally validated positive interaction pairs, covering 22,886 active compounds and 102 protein targets. The dataset undergoes a rigorous preprocessing workflow: all negative samples are removed, and overlaps with the TDC DTI benchmark dataset are eliminated, ensuring that only experimentally confirmed positive interactions are retained. The atlas provides multimodal attribute data, including drug SMILES expressions, protein sequences, and quantitative affinity scores such as IC50, Ki, and Kd, and is standardized using UniProt accession numbers and ChEMBL/DrugBank IDs for identification. The dataset is released in CSV/TSV formats and is primarily used for fair performance evaluation and benchmark testing of drug-target prediction models, providing clean, data-leakage-free training and testing data for computational drug discovery research.
Chemistry

@ZJU
ProteinKG65 is a large-scale knowledge graph dataset for protein science. It aligns protein sequences with functional and ontological knowledge (from Gene Ontology, GO) and is compatible with structured knowledge, sequences, and textual descriptions.
Biology

@ZJU
Amino Acid Knowledge Graph (AAKG) is a domain-specific knowledge graph that systematically represents the biochemical properties of amino acids and their intrinsic relationships.
Biology

@ZJU
OneGraph is a general-purpose knowledge graph constructed with the assistance of large language models. It spans common-sense concepts and relations across the humanities, engineering and technology,
Biology

@ZJU
InstructProteinKG is a protein knowledge graph tailored for protein sequence-text alignment and instruction learning. Primarily extracted from the high-quality
Biology

DREAMwalk
@ZJU
The dataset utilized by DREAMwalk is a multi-layer heterogeneous biomedical knowledge graph. It was not constructed from scratch, but rather formed a foundational network by integrating multiple authoritative public databases (such as DrugBank, DisGeNET, STRING, etc.), and dynamically incorporated semantic information on this basis. Its core innovation lies in utilizing the hierarchical structure of Ontology to calculate semantic similarities among entities of the same type, and incorporating these similarities as new “virtual edges” into the original graph, thereby creating a richer and more biologically semantic-capturing enhanced knowledge graph specifically designed to improve the predictive performance of tasks such as drug repositioning.
Biology

E. coliKG
@ZJU
The KIDS E. coli Antibiotic Resistance Knowledge Graph is a systems biology knowledge graph jointly developed by the Tagkopoulos Laboratory at UC Davis, Zhejiang University, and Fudan University. This graph focuses on antibiotic resistance (AMR) research across the entire genome of Escherichia coli (E. coli), aiming to accelerate the discovery of resistance genes through knowledge integration and automated reasoning.
Biology

OPB
@ZJU
OPB is a reference ontology that applies classical physics and thermodynamics to the dynamics of biological systems. It is designed to encompass multiple structural scales (from atoms to organisms) and various physical fields (fluid dynamics, chemical kinetics, particle diffusion, etc.) encountered in the research and analysis of biological organisms.
Biology

ProteinKG25
@ZJU
ProteinKG25 is a large-scale biomedical knowledge graph (KG) oriented towards understanding protein function, developed by the ZJUNLP team at Zhejiang University. It aims to support the pre-training of knowledge-enhanced protein language models. This graph integrates structured ontological relationships from Gene Ontology (GO) with high-quality protein sequences and annotation information from Swiss-Prot, covering core entities such as gene ontology terms, proteins, and their semantic associations.
Biology

RNA-KG
@ZJU
RNA-KG v2.0 is an RNA-centered heterogeneous biomedical knowledge graph (KG). It not only integrates traditional “entity-relationship” information but also introduces contextual properties. It addresses the common issue of “disconnection from context” in biomedical data, enabling precise description of the specific biological contexts in which RNA interactions occur.
Biology

PhySci
@ZJU
PhySci is an ontology dedicated to scientific publications in physics research. By defining the overall structure of scientific publications, it constructs a knowledge base representation in the field of physics research. Utilizing the principles of Linked Data, it is developed with metadata concepts such as scientific methods, scientific problems, and solutions, achieving an interactive semantic model for heterogeneous data across sub-fields.
Biology

PertKGE
@ZJU
PertKGE is a compound-protein interaction prediction dataset released by the Shanghai Institute of Materia Medica, Chinese Academy of Sciences, and Tsinghua University in 2024. This atlas innovatively constructs a fine-grained knowledge graph that integrates perturbation transcriptomics. By dismantling genes into multiple levels of nucleic acid entities such as DNA, mRNA, lncRNA, and miRNA, and combining compounds, proteins, and transcription factors, it systematically describes the central dogma process of biological systems. The dataset integrates large-scale transcriptome perturbation data such as CMap/LINCS with various public biological network resources, containing approximately 80,000 entities and over 6.8 million triples, including 4.36 million regulatory relationships between compounds and mRNA, and 15,000 physical binding relationships between compounds and proteins. This atlas is primarily applied to new compound target prediction, ligand virtual screening, and drug action mechanism analysis. It effectively mitigates systematic biases in transcriptome data through knowledge graph embedding technology, enhancing the biological interpretability of predictions.
Biology

Cisreg
@ZJU
Cisreg is a gene regulatory knowledge graph dataset integrating chromosome location and function, released by institutions such as the University of Murcia in Spain and the Norwegian University of Science and Technology in 2024. The graph is built based on the Resource Description Framework standard, integrating 25 enhancer databases and 2 topological association domain resources, systematically describing the complex regulatory relationships between entities such as enhancers, topological association domains, genes, transcription factors, and phenotypes. The dataset comprises nearly 1 billion triples, including approximately 683 million pieces of enhancer sequence metadata, 263 million enhancer-target gene associations, and 3.8 million pieces of topological association domain structural information. The graph deeply integrates enhancer sequences, genomic coordinates, and biological functional semantics (such as transcription factor binding and phenotype association), supporting complex regulatory queries that combine spatial position constraints with functional semantics. It is primarily applied to the analysis of long-range gene expression regulatory mechanisms, the assessment of the impact of mutations in non-coding regions, and interoperability research across enhancer databases.
Biology

CKG
@ZJU
CKG is an open-source, large-scale biomedical knowledge graph platform that provides a complete database dump file, containing over 16 million nodes and 220 million relationships. This graph integrates nine biomedical ontologies, 26 public databases, and published experimental research data, unifying vast amounts of heterogeneous biomedical information into a graph structure. Its main application value lies in accelerating proteomics data analysis, supporting clinical decision-making (such as biomarker discovery), and facilitating knowledge discovery. It enables users to uncover hidden associations between entities such as genes, proteins, diseases, and drugs through graph database queries. The dataset is released in the form of a Neo4j database dump file, allowing users to set up a fully functional graph database instance locally for exploration.
Biology

IDP-KG
@ZJU
IDP-KG is an integrated knowledge graph dataset of intrinsically disordered proteins released by the University of Padova, Heriot-Watt University, and the ELIXIR IDP community in 2022. This graph adopts a fully automated ETL process, integrating the Bioschemas annotation data from three professional databases: DisProt, MobiDB, and PED. It unifies entities such as intrinsically disordered proteins, sequence annotations, sequence ranges, and IDPO ontological terms into the RDF standard format. The dataset contains 149 different IDPO codes, with 7,542 disorder annotations and 1,325 protein binding annotations, and fully records the source and traceability information of the data. This graph is primarily used to address the issue of inconsistent identifiers among disordered protein databases, supports cross-database federated queries and in-depth functional analysis, provides high-quality training labels for machine learning models, and serves as a benchmark dataset for the large-scale application of Bioschemas standards in the field of life sciences.
Biology

SpaTalk
@ZJU
SpaTalk is a knowledge graph-based dataset for inferring cell communication, developed by Zhejiang University, specifically designed for spatial transcriptomic data analysis. This graph constructs a three-tier knowledge framework of “ligand-receptor-target”, encompassing both human and mouse species, including 3,324 pairs of human and 2,484 pairs of mouse ligand-receptor pairs, as well as tens of thousands of intracellular signaling pathways. The dataset integrates multiple database resources such as CellTalkDB, KEGG, Reactome, and AnimalTFDB, systematically describing the complete transmission process of cell-to-cell communication signals from extracellular ligands to intracellular target genes through four relationship types: ligand-receptor binding, intracellular interaction, pathway assignment, and transcription factor annotation. This graph is primarily applied to inferring the communication intensity between adjacent cells in spatial transcriptomic data, tracing intracellular signaling pathways, and studying tissue microenvironment, supporting the data analysis of various spatial transcriptomic technologies such as STARmap, Slide-seq, and 10X Visium.
Biology

@ZJU
PorousKG is a knowledge graph of porous materials centered on metal–organic frameworks (MOFs) and extended to covalent organic frameworks (COFs), hydrogen-bonded organic frameworks (HOFs), and cage-like materials.
Materials

Material-KG
@ZJU
Material-Knowledge-Graph is not a directly downloadable “finished” knowledge graph (such as an RDF file or a Neo4j database), but rather an open-source project and accompanying dataset for automatically constructing a knowledge graph of materials science. Its core objective is to utilize a large language model (LLM) to automatically extract structured knowledge from a vast amount of materials science literature, thereby building a knowledge graph encompassing elements such as material names, chemical formulas, properties, and synthesis methods.
Materials

KG-FM
@ZJU
KG-FM is a large-scale framework material knowledge graph constructed by the MontageBai team between 2024 and 2025, specifically designed for metal-organic frameworks, covalent organic frameworks, and hydrogen-bonded organic frameworks. This graph utilizes a large language model to automatically extract structured knowledge from over 100,000 academic papers, constructing a knowledge network comprising 2.53 million nodes and 4.01 million relationships, covering core entity types such as framework materials, synthesis methods, performance indicators, application fields, chemical components, and literature. The graph supports multi-hop reasoning queries through semantic relationships such as has_synthesis_method, exhibits_performance, and used_for. It enables precise Cypher retrieval based on the Neo4j graph database and integrates an intelligent question-answering system based on the Qwen2 large model, providing answers with traceable literature citations. It is primarily applied to rapid retrieval of framework material properties and applications, research trend analysis, hotspot mining, as well as AI-driven material discovery and inverse design.
Materials

MatKG
@ZJU
MatKG is a large-scale, comprehensive Knowledge Graph (KG) for materials science, developed by a research team at the Massachusetts Institute of Technology (MIT). Utilizing advanced Natural Language Processing (NLP) technology, it automatically extracts entities and relationships from over 5 million scientific papers in the field of materials science, constructing a structured knowledge base. This graph aims to address the issue of highly fragmented knowledge in the field of materials science, which is difficult to integrate and utilize, providing a robust data foundation for applications such as material discovery, recommendation systems, and advanced analysis.
Materials

SSC-CoT-KG
@ZJU
The SSC-CoT-KG is a structured knowledge base specifically designed for mathematical reasoning in large language models (LLM). It focuses on formula derivation and logical path retrieval in the fields of trigonometry and geometry. It addresses the common “logical leap” problem encountered by large models in complex mathematical reasoning, where the model may be interrupted in the multi-step derivation process due to forgetting or inability to recall key formulas. The knowledge graph guides the model to construct a complete and coherent reasoning chain by providing precise formula clues for intermediate steps.
Mathematics

OM
@ZJU
OM is a comprehensive unit of measure ontology that provides standardized representations for units, dimensions, and related concepts. It supports scientific computation, data exchange, and semantic integration, and is particularly suitable for scenarios requiring precise unit conversion and dimensional analysis.
Physics
